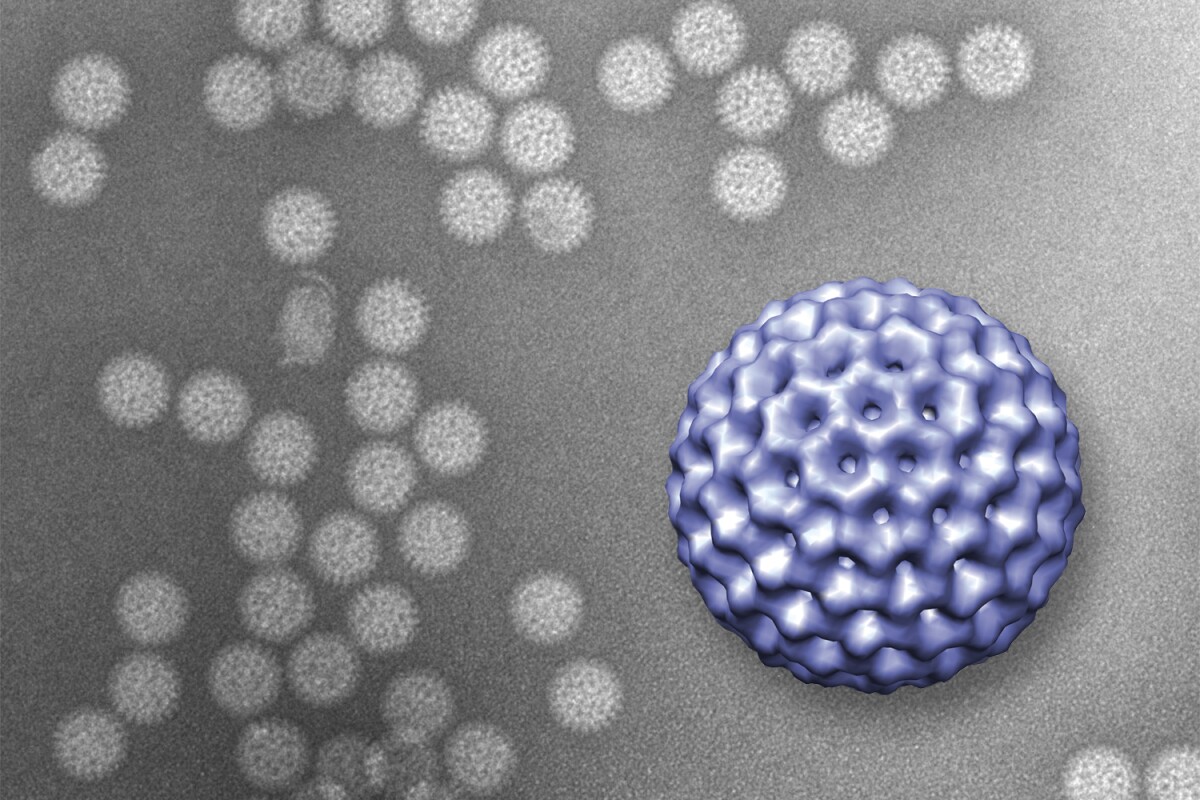
Traditionally, in order to view tiny biological structures such as viruses, they must first be removed from their natural habitats and frozen. While this certainly keeps them still for the microscope, it greatly limits what we can learn about them – it’s comparable to an ichthyologist only being able to study dead fish in a lab, instead of observing live ones in the ocean. Now, however, researchers at the Virginia Tech Carilion Research Institute have devised a technique for observing live viruses in a liquid environment. It could have huge implications for the development of treatments for viral infections.
The team was led by assistant professors Deborah Kelly and Sarah McDonald.
They started by taking two silicon-nitride microchips that had windows etched into their centers, and then pressing them together until a gap measuring just 150 nanometers remained between them. That space was then filled with a liquid similar to that in which the rotavirus (the most common cause of severe diarrhea in infants and children) is normally found. The inside surfaces of the windows were then coated with antibodies.
When a rotavirus was subsequently injected into the chamber of liquid, the antibodies latched onto it and held it in place between the windows. A transmission electron microscope was then used to image the virus, producing results similar in quality to those obtained using a conventional freezing approach – in this case, however, the virus was still alive and in its natural medium.
The scientists now hope to use the technique to actually observe the rotavirus in action, to better understand how it uses an infiltrated host cell to create more viruses. By knowing more about this process, it’s possible that new approaches could be developed for fighting the virus, potentially saving countless young lives in the developing world.
Along with viruses, other microscopic structures could also be viewed using the technology – different fluids and “biological tethers” (the role played by the antibodies) simply might have to be used, depending on the target.
“What’s missing in the field of structural biology right now is dynamics – how things move in time,” said McDonald. “Debbie is developing technologies to bridge that gap, because that’s clearly the next big breakthrough that structural biology needs.”
A paper on the research was recently published in the journal Lab on a Chip.