Researchers have identified 317 million unique gene clusters belonging to oceanic microbes. Creating the world’s largest open-source catalog representing only the ‘tip of the iceberg’ in marine metagenomics, the library offers a tool for exploring how these genetic resources could be used in medicine, energy, food and other industries.
The ocean’s microbiome is a vast, highly diverse gene pool with complex metabolic capabilities. The global ocean genome has already proven to be a major resource for science, particularly in the area of health. For example, the green fluorescence protein that was first isolated from jellyfish that’s now widely used in medical imaging diagnostics and the bacteria living around hydrothermal vents that were the source of the polymerases used in PCR tests to detect SARS-CoV-2. But there are many, many more genes to be discovered.
Metagenomics, the study of genetic material taken directly from environmental or clinical samples, allows gene function to be matched with the organism to which the gene belongs. Analyzing the genetic makeup of the millions of oceanic microbes is a huge undertaking. Thankfully, the rise of AI and improvements in computational power have enabled massive-scale metagenomic analysis.
Now, researchers from the King Abdullah University of Science and Technology (KAUST) have collaborated with the Institute of Marine Sciences at the Spanish National Research Council (CSIC) to analyze and catalog massive amounts of genetic information about the microbes that dwell in our oceans.
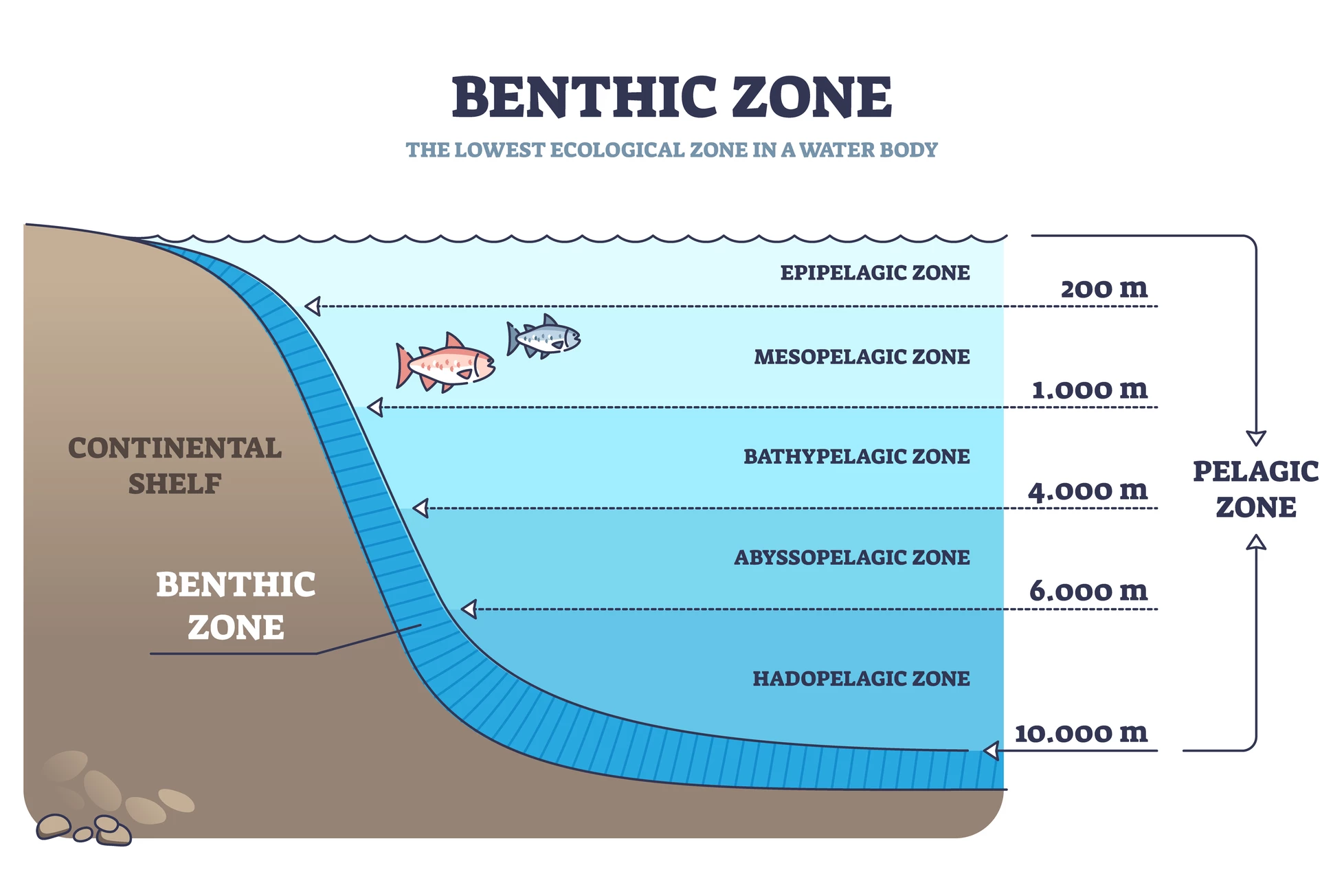
The researchers used the KAUST Metagenomic Analytical Platform (KMAP), invented in 2021, to analyze 2,102 ocean samples. Most (78.5%) samples were collected in the upper ocean (zero to 200 m/656 ft); 7.2% were from the mesopelagic zone (200 m/656 ft to 1,000 m/3,281 ft); and 10.2% from the dark ocean, at depths below 1,000 m/3,281 ft.
Their DNA sequencing analyses identified 317.5 million unique gene clusters, which they used to create the KMAP Global Ocean Gene Catalog 1.0, the world’s largest open-source catalog of marine microbes that matches a microbe with gene function, geographic location and habitat type. Beyond advancing our understanding of the ocean microbiome and its metabolic capabilities, the information provided enables scientists to track global warming, pollution and overall ocean health, as well as providing a tool to explore the potential uses of novel genes in medicine, energy, food, and other industries.
“Scientists can access the catalog remotely to investigate how different ocean ecosystems work, track the impact of pollution and global warming, and search for biotechnology applications such as new antibiotics or new ways to break down plastics,” said Carlos Duarte, corresponding author of the study. “The acceleration of AI we are currently experiencing is likely to play a major role in identifying genes of biotechnology interest contained within the massive catalog we are releasing.”

Interestingly, fungi represented over 50% of the distinct gene clusters identified in the mesopelagic zone, highlighting the contribution of fungi to microbial diversity. And, while 95.9% of the samples were from the pelagic zone, the open, free water away from the shore, 4.1% were from the benthic zone, the ocean floor. Benthic microbes are pivotal to oceanic biogeochemical cycles, the turnover of vital elements like carbon, nitrogen, and sulfur facilitated by interactions between living (biotic) and non-living (abiotic) parts of the biosphere. Gathering information about these microbes provided invaluable information about how marine ecosystems are adapting to an environment that’s constantly changing due to natural and anthropogenic causes.
“Our analysis highlights the need to continue sampling the oceans, focusing on areas that are understudied, such as the deep sea and ocean floor,” said Elisa Liaolo, the study’s lead author.
While 317.5 million probably sounds like a lot of gene clusters, the researchers understand they still have their work cut out for them.
“Impressive as it is, the 317 million gene groups documented in the Ocean Gene Catalog 1.0 likely represents the tip of the iceberg of the massive library of functional capacities the long evolutionary history of life in the ocean has accrued,” Duarte said. “Further projects focused on sampling and massive sequencing of understudied habitats in the ocean, including organisms such as corals and seagrass, not included in the study, which are known to host large numbers of microbial species, will likely reveal many times the number of genes included in this initial gene catalog.”
The study was published in the journal Frontiers in Science.
Source: KAUST